3-3.2.2 Real Time PCR or Quantitative PCR (qPCR):
Real-time or Quantitative PCR (qPCR) uses the linearity of DNA amplification to quantify absolute or relative amounts of target sequence in a sample. With the help of a fluorescent reporter, the amount of generated DNA can be measured. In qPCR, DNA amplification is monitored at each cycle of PCR. When the DNA is getting amplified logarithmically at each cycle, the amount of fluorescenceincreases over the basal level. The thermal cycle at which the signal exceeds the fluorescence detection threshold is known as the Threshold cycle (CT) or crossing point. A standard curve of log concentration against CT can be made by making use of multiple dilutions of a known amount of standard DNA. The quantity of DNA or cDNA in an unknown sample can thus be determined from its CT value.
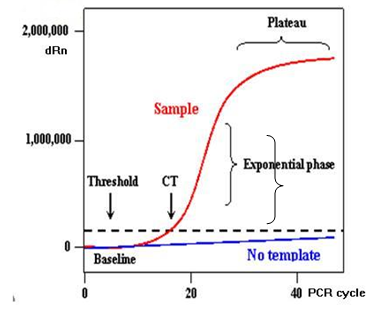
Fig 3-3.2.2: A Representation of Real Time PCR Plot (Obtained from NCBI Probe)
(Baseline is defined as PCR cycles in which a reporter fluorescent signal is accumulating but is beneath the limits of detection of the instrument. dRn is an increment of fluorescent signal at each time point.)
There are two detection chemistries used for q PCR:
a) Use of an intercalating dye like the SYBR® Green I dye which incorporates between the base pairs of DNA. This detection method is suitable when the PCR reaction generates a specific product, as the dye is capable of intercalating into any double-stranded DNA product.
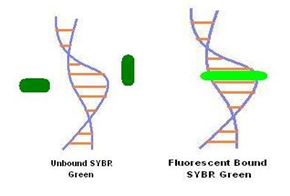
Fig 3-3.2.2.1: SYBR Green I Detection Mechanism
(The dye intercalates with double-stranded DNA in the reaction. In the bound state, SYBR Green I exhibits 1000 fold more fluorescence than the unbound state. The fluorescence signal increases in proportionwith the increase in amplified DNA)