3-5.6.1 Labeling 3' Termini of dsDNA with recessed 3' termini or blunt-ended:
Klenow fragment of E.coli DNA polymerase I is used to incorporate 32P dNTPs into a recessed 3' terminus generated by restriction digestion. The choice of 32P dNTP for labeling reaction depends on sequence of protruding 5' termini of DNA. End fragments created by restriction endonucleases can be labeled with 32P dATP.
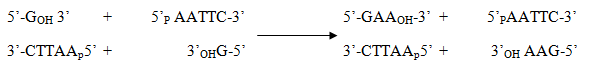
The terminal nucleotide of blunt end DNA fragment can be replaced by weak 3'→5' exonuclease and strong polymerase activity of klenow fragment.
3-5.6.2 Labeling 3'termini of dsDNA with protruding 3'termini:
Bacteriophage T4 DNA polymerase have strong 3'→5' exonuclease activity than of klenow fragment. It is used to digest 3'-protruding termini and then continues at slower pace to remove 3' nucleotides from ds DNA. At higher concentration of dNTPs, radiolabelled dNTP is added by polymerase activity.
Alternatively, protruding 3'termini can be labeled by calf thymus terminal transferase to catalyze the transfer of 32P dideoxy ATP. Because it does not carry 3'-OH group, no additional nucleotides can be added to protruding 3'end.
3-5.6.3 Labeling RNA probes:
RNA probes yield stronger signals in hybridization reactions than DNA probes of equal specific activity which may be due to innately higher stability of hybrid involving RNA. They are the probes of choice when analyzing transcripts of mammalian genes.
RNA probes are most easily achieved by in vitro transcription in which insert DNA is cloned adjacent to multiple cloning sites in suitable plasmid expression vector. This expression vector has a phage promoter sequence which is recognized by corresponding phage RNA polymerase. In the given plasmid vector pSP64, bacteriophage SP6 promoter sequence is recognized by SP6 RNA polymerase which starts transcription from a particular initiation site after which insert has been cloned. Highly specific labeled RNA probes can be generated using radiolabeled dNTPs.