3-5.5.1.2 Random primed DNA labeling:
This method known as oligo-labeling is based upon hybridization of a mixture of all possible hexanucleotides. The template DNA is initially denatured and then cooled slowly so as to allow random hexanucleotides to bind at complementary sequences at which extension takes place through PCR.
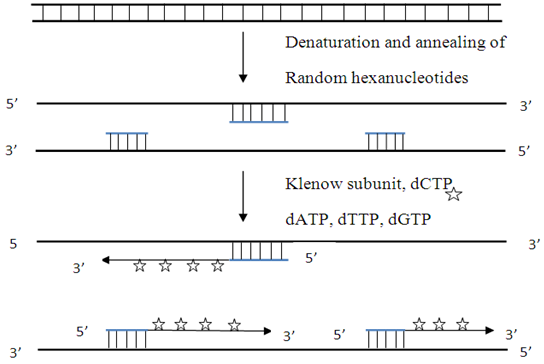
Fig 3-5.5.1.2: Random primed DNA labeling
The synthesis of new complementary DNA strands is primed by bound hexanucleotides and catalyzed by Klenow subunit of DNA polymerase I (contains the polymerase activity in the absence of associated 5'→3' exonuclease activities, sometimes 3'→5' exonuclease activity also removed for the reaction).This method produces labeled DNAs of high specific activity. Since binding of primer to template DNA is random and primer represents all possible sequence combinations,uniform labeling of DNA occurs.
The length of random primers is crucial. Primers shorter than 6 bases are very poor primers, whereas those longer than 7 have higher tendency to self–anneal and cause primer duplex. Thus, probe should be either 6 or 7 nucleotides in length.
Random priming is inherently simpler than nick translation because the requirements for two nuclease activities are eliminated. The probes generated by random priming method are more homogenous in size and behave more reproducibly in hybridization reactions. Average size of probe cannot be controlled with great accuracy in nick translation method.
Template DNA used preferably should not be closed, circular ds DNA because they are inefficient templates. Shorter templates generate probes of low specific activity that may not hybridize under stringent conditions.