3-4.3.1 Procedure:
The northern blotting involves the following steps:
1. Total RNA is extracted from a homogenized tissue sample or cells. Further eukaryotic mRNA can then be isolated by using of oligo (dT) cellulose chromatography to isolate only those RNAs by making use of a poly A tail.
2. The isolated RNA is then separated by gel electrophoresis.
3. The RNA samples separated on the basis of size are transferred to a nylon membrane employing a capillary or vacuum based system for blotting.
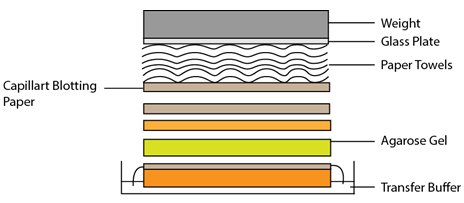
Fig 3-4.3.1: Setup for Northern blotting
4. Similar to Southern blotting, the membrane filter is revealed to a labeled DNA probe that is complementary to the gene of interest and binds.
5. The labeled filter is then subjected to autoradiography for detection.
The net amount of a specific RNA in a sample can be estimated by using Northern blot. This technique is widely used for comparing the amounts of a particular mRNA in cells under different conditions. The separation of RNA samples is often done on agarose gels containing formaldehyde as a denaturing agent as it limits the RNA to form secondary structure.
3-4.3.2 Analysis of Northern Blot:
RNA extract is electrophoresed in an agarose gel, using a denaturing electrophoresis buffer (containing formaldehyde) to ensure that the RNAs do not form inter- or intra-molecular base pairs, as base pairing would affect the rate at which the molecules migrate through the gel. After electrophoresis, the gel is blotted onto a nylon or nitrocellulose membrane, and hybridized with a labeled probe. If the probe is a cloned gene, the band that appears in the autoradiograph is the transcript of that gene. The size of the transcript can be determined from its position within the gel, and if RNA from different tissues is run in different lanes of the gel, then the possibility of differentially expressed gene can be examined.