3-4.1 Introduction:
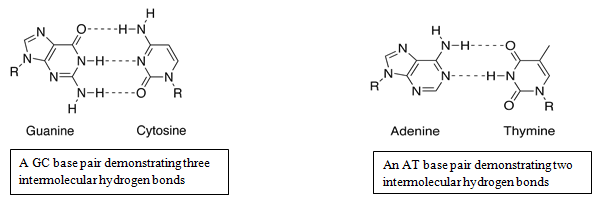
Nucleic acid hybridization is a basic technique in molecular biology which takes advantage of the ability of individual single-stranded nucleic acid molecules to form double-stranded molecules. According to Watson-Crick base pairing, adenine binds with thymine and guanine binds with cytosine by hydrogen bonding.
3-4.2 Southern hybridization:
The basic principle behind the southern hybridization is the nucleic acid hybridization. Southern hybridization commonly known as southern blot is a technique employed for detection of a specific DNA sequence in DNA samples that are complementary to a given RNA or DNA sequence. It was the first blotting technique to be devised, named after its pioneer E.M Southern, a British biologist. Southern blotting involves separation of restricted DNA fragments by electrophoresis and then transferred to a nitrocellulose or a nylon membrane, followed by detection of the fragment using probe hybridization.
Separated by electrophoresis is transferred from gel to a membrane which in turn is used as a substrate for hybridization analysis employing labeled DNA or RNA probes specific to target fragments in the blotted DNA. Southern hybridization helps to detect specific fragment against a background of many other restriction fragments. Southern blotting is a technique which is used to confirm the identity of a cloned fragment or for recognition of a sub-fragment of interest from within the cloned DNA, or a genomic DNA. Southern blotting is a prerequisite to techniques such as restriction fragment length polymorphism (RFLP) analysis.