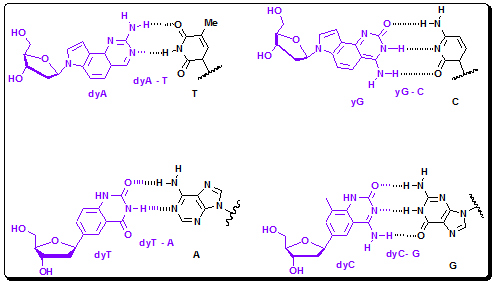
Figure 4.36b: Structures and H-bonding pattern of yDNA bases.
In a homologous way they again expanded the size of the bases by Inclusion of another benzene ring to the pairs. This would yield 4.8 Å of expansion, giving hypothetical xxDNA and yyDNA ("double-wide DNA") designs (Figure 4.37). The possible self-assembly of the double-wide DNA helix establishes a new limit for the size of information-encoding, DNA-like molecules, and the fluorescence of yyDNA bases suggests uses as reporters in monomeric and oligomeric forms.
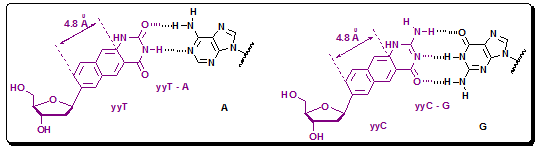
Figure 4.37: The base pairs of doublewide DNA
Another type of extended DNA pair was recently reported by Inouye, based on the acetylene-linked C-nucleosides bearing pyrimidine heterocycles capable of forming Watson−Crick-like H-bonds (iG*−iC* and A*−T*) (Figure 4.38).
Two interesting pairs of extended imidazolopyridopyrimidines and naphthyridine C-nucleosides (ImON/NaNO and ImNO/NaON) were prepared by Minakawa and Matsuda (Figure 4.38). Their four-H-bond interactions ensure extra stable duplexes and moderately selective recognition and incorporation by Klenow fragment polymerase.
Hirao et al. have reported the nucleobases y and v (Figure 4.38) and showed that dyTP is incorporated into DNA template opposite to dx more efficiently than any natural substrate. The dyTP was incorporated into the template opposite to ds with higher efficiency and a 3-fold higher selectivity than opposite to dx. The efficiency and fidelity of y−v pairing were as high as in natural base-pairs. The y−s pair was used for in vitro incorporation of chlorotyrosine into a protein proving a new genetic codon system.
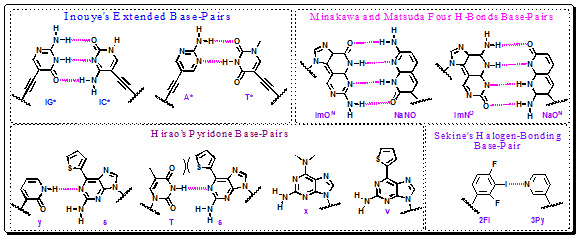
Figure 4.38: Few more unnatural base-pairs that are paired via H-bonding
An interesting alternative to H-bonds was studied by Sekine, who endeavored to employ halogen bonding for specific base-pairing of artificial nucleosides. Out of several halogenated benzene and pyridine pairs, the most stable and selective binding was found for difluoroiodobenzene (2FI) and pyridine (3Py) pair, but no evidence for the halogen bond was provided (Figure 4.38).
It is quite clear that unnatural base-pairs that are paired via H-bonding patterns offer promising approaches to expanding the genetic alphabet. However, there are some inherent limitations to their selective replication in DNA, due to the population of minor tautomeric forms of the unnatural bases that form mispairs.