4.19.4.1.3.2. Cytosine Analogue
Benzo[g]quinazoline based cytosine base analog: A 2'-O-Me ribonucleoside derivative of 4-amino-1Hbenzo[g]quinazoline-2-one has also been synthesized based on the same heterocyclic benzo[g]quinazoline design and used as a novel fluorescent cytosine base analog probe. This cytosine base analog exhibits a fluorescence emission centered at 456 nm, characterized by four major excitation maxima (250, 300, 320 and 370 nm) and a fluorescence quantum yield ΦF = 0.62 at pH= 7.1. The fluorescence emission of this probe shifted from 456 to 492 nm when pH was decreased from 7.1 to 2.1. The pKa (4.0) of the probe was close to that of cytosine (4.17). This probe was used to detect the protonation state of base triplets in triple stranded structures.
Tricyclic cytosine (tC): Norden and colleagues have described a new cytosine base analog, 3,5-diaza-4-oxophenothiazine or tricyclic cytosine (tC), which can form a specific base pair with guanine (Figure 4.34). Like the benzo[g]quinazoline base analogs, this base maintains its relatively high quantum yield (ΦF= 0.20) even after incorporation into single and double stranded oligonucleotides, like artificial peptide nucleic acid (PNA) biopolymers and RNA-DNA duplexes.
Elaboration of the tC(O) scaffold can yield a nitroxide spin labeled compound that may be used for EPR measurements and the “G-clamp” which has increased binding affinity to guanine (A, C, D) (Figure 4.34).
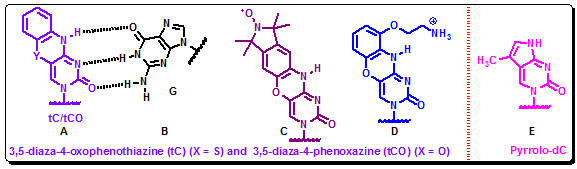
Figure 4.34: 3,5-diaza-4-oxophenothiazine (tC) (X = S) and 3,5-diaza-4-phenoxazine (tCO) (X = O) in hybridization with guanine (A, C, D). The structure of pyrrolo-dC (E).
Pyrrolo-dC: cytosine Analog: In another study, a new highly fluorescent analog of cytosine, pyrrolo-dC, (Figure 4.34) has been introduced to characterize the transcription bubble in elongation complexes of T7 RNA polymerase. Pyrrolo-dC has excitation and emission maxima at 350 nm and 460 nm, respectively, which, like the previously described analogs, allows selective excitation in the presence of native nucleic acid bases and proteins. This base analog can pair with guanine and, like 2-AP, pteridine and hydrocarbon base analogs, shows significant quenching of fluorescence when incorporated into single and double stranded DNA. The quenching can be used to monitor local melting of the G:C base pairs in a DNA helix and can serve as a complementary probe to 2-AP, which reports on melting of AT base pairs.
4.19.4.1.4. Xanthosine Analogue
In addition to these cytosine and thymine analogs, the synthesis of the fluorescent analog of the rare base xanthosine (5-aza-7-deazaxanthine) has also been reported (Figure 4.35). This base was designed based on the supposition that the rare tRNA constituent wyosine carries the 5-aza-7-deazapurine substructure, and it is this structure that makes the base fluorescent. This base analog can be excited at 250 nm and displays two emission maxima at 410 and 580 nm.
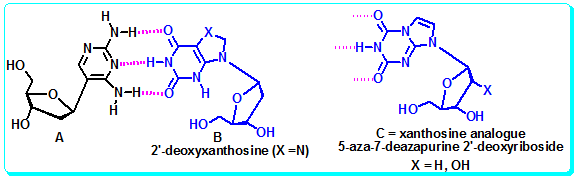
Figure 4.35: The Xanthosine analogue
4.19.4.2 Extended Base-Pairs/Base-Pairs Based on Four H-Bonds/Halogen-Bonding Base-Pair:
Most, recently Kool and coworkers are interested in designing much more size expanded nucleobases for new genetic bases having interesting fluorescent properties. Thus, they reported the synthesis, and properties of DNA-like molecules in which the base pairs are expanded by benzo homologation. The resulting size-expanded genetic helices are called xDNA ("expanded DNA") and yDNA ("wide DNA"). The large component bases are fluorescent, and they display high stacking affinity. When singly substituted into natural DNA, they are destabilizing because the benzo-expanded base pair size is too large for the natural helix. However, when all base pairs are expanded, xDNA and yDNA form highly stable, sequence-selective double helices. The size-expanded DNAs are candidates for components of new, functioning genetic systems. In addition, the fluorescence of expanded DNA bases makes them potentially useful in probing nucleic acids (Figure 4.36a-b).
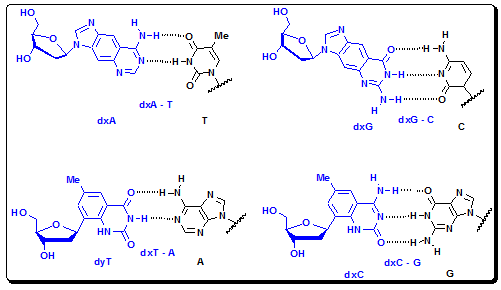
Figure 4.36a: Structures and H-bonding pattern of xDNA bases.