4.19. Expanding the Genetic Alphabets: Background
DNA is an essential biomolecule which is responsible for encoding the complex information necessary for life. Specific pairing of dA with dT and dC with dG in duplex DNA and during polymerase-mediated replication is the basis of the genetic alphabet, itself the basis of the genetic code. However, there is no reason to assume that the requirements for duplex stability and replication must limit the genetic alphabet to only two base pairs. In addition to enabling a wide variety of biotechnology applications, an expanded genetic alphabet would enable the encoding of additional information for both in vitro and in vivo applications. Expansion of the genetic alphabet to include a third base pair, formed between two identical or different unnatural nucleotides, referred to as self-pairs and hetero pairs, respectively, would expand the informational and functional potential of DNA such as site directed oligonucleotide labeling and in vitro selections with oligonucleotides bearing increased chemical diversity. The use of the efficient new base pair/pairs to drive the synthesis of proteins with unnatural base pairs is also a current exciting research area to consider. Thus, translation of an expanded genetic alphabet into an expanded genetic code, provocatively, even lead to the assembly of such a system within a living cell, potentially creating a semi-synthetic organism and life with increased diversity.
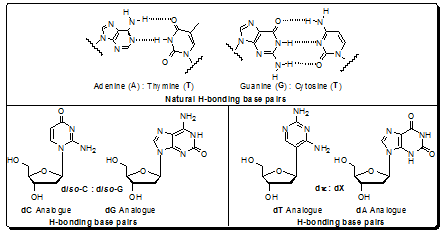
Figure 4.26: Presentation of H-bonding base pairs among natural nucleosides and analogues
After Benner first popularized the idea, an increasing amount of interest has resulted in the acceleration of progress towards this goal. Expanding the genetic alphabet requires an unnatural base pair with inter base interactions, of whatever sort, that confer stability on a DNA duplex and that is replicated by a DNA polymerase. Specifically, each unnatural triphosphate must be efficiently and selectively incorporated opposite its partner in the template to form a stable base pair. Conversely, no natural substrate should be inserted opposite the unnatural nucleotide in the template with high efficiency. Also, continued synthesis past the unnatural base pair must be efficient. Thus, the efforts toward developing a third base pair have focused on nucleobase analogues designed to pair via orthogonal hydrogen bonding (H-bonding, Figure 1), based on work of the Benner group, and more recently on predominantly non-H-bonding (Figure 2) analogues that pair via hydrophobic interactions, based on work of the Kool group.
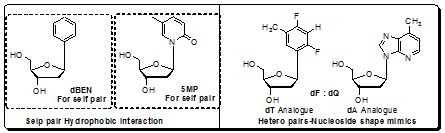
Figure 4.27: Presentation of non H-bonding base pairs among nucleoside base analogues
However, the unnatural base pairs so far have been reported has several shortcomings, including tautomerization of iso-G and poor recognition of iso-C by RNA polymerases; these shortcomings pose difficulties for mRNA preparation. Novel, hydrophobic base pairs have been developed recently, but their use in transcription is still under investigation. Thus there is a need to develop conceptually new and novel base analogues which will be recognized by both replication and transcription process with high efficiency.
4.19.1. Synthesis and Application of Unnatural Nucleosides
Virtually all modern molecular biology techniques require the amplification of DNA by PCR. Thus, an unnatural base pair that is compatible with both PCR amplification and in vitro transcription, expanding the genetic alphabet, at least in a test tube, from just two base pairs to three. It will be worthwhile to design such unnatural pair for which both efficient PCR amplification and transcription by RNA polymerase would efficient. In addition to the incorporation of new base pairs into DNA or RNA, the unnatural base that would allow site-specific labeling of RNA will be of great importance for a variety of biotechnology applications.1 For example, the ability to site-selectively modify a DNA or an RNA molecule with a fluorophore attached to the unnatural base pair should facilitate applications in both cell biology and biophysics. The third base pair should also be useful for in vitro selection methodologies that have already produced DNA and RNA molecules with desired properties, such as selective recognition of other molecules (aptamers) or catalysis (ribozymes). Increasing diversity of these modified DNA and RNA molecules, promises their enhanced widespread potential applications in biomedical sciences such as drug candidates.