The following is a list of experimentally validated riboswitches, organized by ligand.
- Cobalamin riboswitch → regulate cobalamin biosynthesis and transport of cobalamin and similar metabolites, and other genes.
- cyclic di-GMP riboswitches →bind the signaling molecule cyclic di-GMP in order to regulate a variety of genes controlled by this second messenger.
- FMN riboswitch → regulate riboflavin biosynthesis and transport.
- glmS riboswitch → is a ribozyme that cleaves itself.
- Glutamine riboswitches → bind glutamine to regulate genes involved in glutamine and nitrogen metabolism, as well as short peptides of unknown function.
- Glycine riboswitch → binds glycine to regulate glycine metabolism genes, including the use of glycine as an energy source.
- Lysine riboswitch → binds lysine to regulate lysine biosynthesis, catabolism and transport.
- PreQ1 riboswitches → bind pre-queuosine1, to regulate genes involved in the synthesis or transport of this precursor to queuosine.
- Purine riboswitches → binds purines to regulate purine metabolism and transport.
- SAH riboswitches → bind S-adenosylhomocysteine to regulate genes involved in recycling this metabolite that is produced when S-adenosylmethionine is used in methylation reactions.
- SAM riboswitches →bind S-adenosyl methionine (SAM) to regulate methionine and SAM biosynthesis and transport. Three distinct SAM riboswitches are known: SAM-I (originally called S-box), SAM-II and the SMK box riboswitch.
- SAM-SAH riboswitches →bind both SAM and SAH with similar affinities. Since they are always found in a position to regulate genes encoding methionine adenosyltransferase, it was proposed that only their binding to SAM is physiologically relevant.
- Tetrahydrofolate riboswitches → bind tetrahydrofolate to regulate synthesis and transport genes.
- TPP riboswitches → binds thiamin pyrophosphate (TPP) to regulate thiamin biosynthesis and transport, as well as transport of similar metabolites. It is the only riboswitch found so far in eukaryotes.
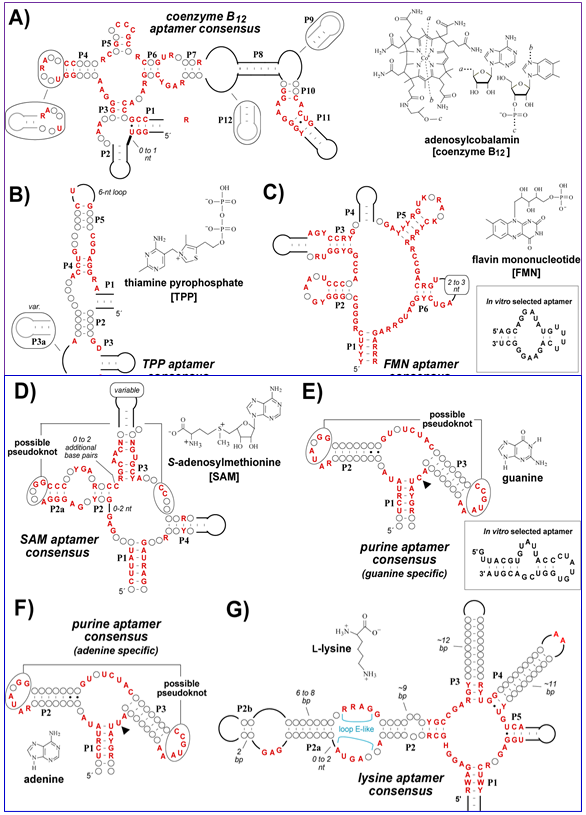
Figure 4.25a: Some riboswitches.
4.18.4.8. Riboswitches and the RNA World hypothesis
Riboswitches demonstrate that naturally occurring RNA can bind small molecules specifically, a capability that many previously believed was the domain of proteins or artificially constructed RNAs called aptamers. The existence of riboswitches in all domains of life therefore adds some support to the RNA world hypothesis, which holds that life originally existed using only RNA, and proteins came later; this hypothesis requires that all critical functions performed by proteins (including small molecule binding) could be performed by RNA. It has been suggested that some riboswitches might represent ancient regulatory systems, or even remnants of RNA-world ribozymes whose binding domains are conserved.
4.18.4.9. Riboswitches as antibiotic targets
Riboswitches could be a target for novel antibiotics. Indeed, some antibiotics whose mechanism of action was unknown for decades have been shown to operate by targeting riboswitches. For example, when the antibiotic pyrithiamine enters the cell, it is metabolized into pyrithiamine pyrophosphate. Pyrithiamine pyrophosphate has been shown to bind and activate the TPP riboswitch, causing the cell to cease the synthesis and import of TPP. Because pyrithiamine pyrophosphate does not substitute for TPP as a coenzyme, the cell dies.