4.18.3.2. Applications of in vivo siRNA delivery in disease models
- RNAi technology is proving to be useful to analyze quickly the functions of a number of genes in a wide variety of organisms.
- RNAi technology has been successfully applied to identify genes with essential roles in biochemical signaling cascades, embryonic development, and other basic cellular process.
- In plants, gene knockdown-related functional studies are being carried out efficiently when transgenes are present in the form of hairpin (or RNAi) constructs. Plant endotoxins could also be removed if the toxin biosynthesis genes are targeted with the RNAi constructs, like theobromine synthase of the coffee plant was knocked down with the hairpin construct of the transgene, leading to the production of decaffeinated coffee plants.
- RNAi may facilitate drug screening and development by identifying genes that can confer drug resistance or genes whose mutant phenotypes are ameliorated by drug treatment, providing information about the modes of action of novel compounds.
- It may also be a method of choice to study the simultaneous functions of a number of analogous genes in organisms in which redundancy exists with respect to a particular function, because many of these genes can be silenced simultaneously.
- siRNAs have been shown to inhibit infection by human immunodeficiency virus, poliovirus, and hepatitis C virus in cultured cell lines. siRNAs can successfully be used to silence genes expressed in respiratory syncytial virus, an RNA virus that causes severe respiratory disease in neonates and infants. siRNA treatment has also been shown to reduce the expression of the BCR-ABL oncoprotein in leukemia and lymphoma cell lines, leading to apoptosis in these cells. With respect to future medical applications, siRNA-based therapy seems to have a great potential to combat carcinomas, myeloma, and cancer caused by over expression of an oncoprotein or generation of an oncoprotein by chromosomal translocation and point mutations.
- Medicine: RNA interference also used in therapy. It is difficult to introduce long dsRNA strands into mammalian cells due to the interferon response; the use of short interfering RNA mimics has been used for medical purpose. Among the first applications to reach clinical trials were in the treatment of macular degeneration and respiratory syncytial virus. RNAi has also been shown to be effective in the reversal of induced liver failure in mouse models.
- Other probable clinical uses: (i) antiviral therapies, like the inhibition of viral gene expression in cancerous cells, (ii) in co-receptors for HIV, (iii) in silencing of hepatitis A and hepatitis B genes (iv) inhibition of measles viral replication, (v) in treatments for neurodegenerative diseases have also been proposed, with particular attention being paid to the polyglutamine diseases such as Huntington's disease, (vi) gene therapy, (vii) in biotechnology.
4.18.4. Riboswitches – mRNA Controls Itself
4.18.4.1. Introduction
A riboswitch is a part of an mRNA molecule that can directly bind a small target molecule, and whose binding of the target affects the gene's activity. Thus, an mRNA that contains a riboswitch is directly involved in regulating its own activity, in response to the concentrations of its target molecule. The discovery that modern organisms use RNA to bind small molecules, and discriminate against closely related analogs, significantly expanded the known natural capabilities of RNA beyond its ability to code for proteins or to bind other RNA or protein macromolecules.
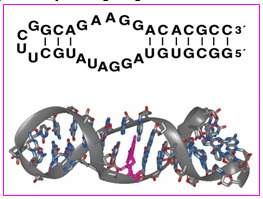
4.18.4. 2. RNA aptamers
|
|
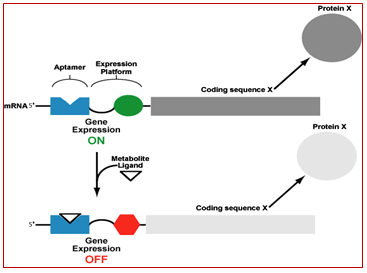
4.18.4. 3. Natural aptamers are the basis of riboswitches:
|
|