4.18.3. RNA Interference (RNAi)
4.18.3.1. Introduction
- Definition: Normal cells contain double-stranded DNA and single-stranded RNA but not double-stranded RNA (dsRNA).
- RNA interference (RNAi), is a technique in which exogenous, double-stranded RNAs (dsRNAs) that are complimentary to known mRNAs, are introduced into a cell to specifically destroy that particular mRNA, thereby diminishing or abolishing gene expression.
- This technology considerably bolsters functional genomics to aid in the identification of novel genes involved in disease processes and thus can be used for medicament and for delivery as therapeutics.
- RNA interference was known by other names, including co-suppression, post transcriptional gene silencing and quelling. Only after these apparently unrelated processes were fully understood did it become clear that they all described the RNAi phenomenon.
- Nomination of RNAi as the “Breakthrough of the year 2002” by the Journal Science prompted biologists to overhaul their vision of the cell and its evolution, and discovery of RNAi was awarded Nobel Prize to Andrew Z. Fire and Craig C. Mello in 2006 in Physiology or Medicine for their work on RNA interference in the nematode worm C. elegans, which they published in 1998.
4.18.3.1. Effectors RNA molecules
RNAi pathways are guided by small RNAs that include:
- SiRNA: Small interfering RNA (siRNA), sometimes known as short interfering RNA or silencing RNA, is a class of 20-25 nucleotide-long double-stranded RNA molecules.
SiRNAs can also be exogenously (artificially) introduced into cells by various transfection methods to bring about the specific knockdown of a gene of interest. - miRNA: microRNAs (miRNA) are single-stranded RNA molecules of about 21–23 nucleotides in length, which regulate gene expression. miRNAs are encoded by genes from whose DNA they are transcribed but miRNAs are not translated into protein (non-coding RNA); instead each primary transcript (a pri-miRNA) is processed into a short stem-loop structure called a pre-miRNA and finally into a functional miRNA. Mature miRNA molecules are partially complementary to one or more messenger RNA (mRNA) molecules, and their main function is to down-regulate gene expression.
- shRNA: A small hairpin RNA or short hairpin RNA (shRNA) is a sequence of RNA that makes a tight hairpin turn that can be used to silence gene expression via RNA interference. shRNA uses a vector introduced into cells and utilizes the U6 promoter to ensure that the shRNA is always expressed. This vector is usually passed on to daughter cells, allowing the gene silencing to be inherited. The shRNA hairpin structure is cleaved by the cellular machinery into siRNA, which is then bound to the RNA-induced silencing complex (RISC). This complex binds to and cleaves mRNAs which match the siRNA that is bound to it.
- Others: In addition to miRNAs and siRNAs, other innate RNAi effectors have been identified. One class of these is the Piwi-interacting RNAs (piRNAs). piRNAs seem to be uniquely expressed in the mammalian germline, particularly in the testes. The functional role of piRNAs is currently unclear, but a role in spermatogenesis is likely.
A number of other small RNAs associated with RNAi have been identified in different species, including trans-activating siRNAs (tasiRNAs), studied in plants and nematodes, and small scan RNAs (ScnRNAs), found in Tetrahymena.
4.18.3.1. Mechanism of RNA interference (RNAi)
Long double-stranded RNAs (dsRNAs; >200 nt) can be used to silence the expression of target genes in a variety of organisms and cell types (e.g., worms, fruit flies, and plants). Upon introduction, the long dsRNAs enter a cellular pathway that is commonly referred to as the RNA interference (RNAi) pathway. First, the dsRNAs get processed into 20-25 nucleotide (nt) small interfering RNAs (siRNAs) by an RNase III-like enzyme called Dicer (initiation step). Then, the siRNAs assemble into endoribonuclease-containing complexes known as RNA-induced silencing complexes (RISCs), unwinding in the process. The siRNA strands subsequently guide the RISCs to complementary RNA molecules, where they cleave and destroy the cognate RNA (effecter step). Cleavage of cognate RNA takes place near the middle of the region bound by the siRNA strand.
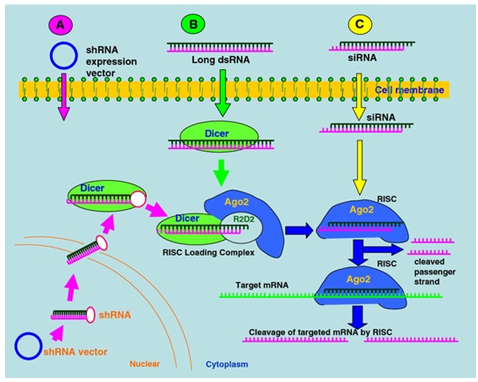
Figure 4.24: Mechanism of RNA interference (RNAi).
In mammalian cells, introduction of long dsRNA (>30 nt) initiates a potent antiviral response, exemplified by nonspecific inhibition of protein synthesis and RNA degradation. The mammalian antiviral response can be bypassed, however, by the introduction or expression of siRNAs.
Natural role of RNAi is defending against RNA viruses
- RNA viruses replicate via dsRNA intermediate (replicative form)
- Applies to RNA virus whether contains ssRNA or dsRNA
- dsRNA is seen as a sign of infection and triggers anti-viral response
RNA interference found in:
- Most eukaryotes - protozoa, invertebrates
- Mammals - weaker
- Plants – especially effective
- NOT found in prokaryotes
- [In bacteria, ribonuclease III rapidly degrades dsRNA molecules]