CD of proteins and peptides
Proteins are the linear polymers made up of 20 amino acids, 19 out of which (except glycine) are chiral. This chirality is also reflected in the secondary structures the proteins/peptides adopt. Far-UV CD spectra of proteins are typically recorded from 190 – 250 nm. Peptide bond is the major chromophore in this region and the relative orientations of the peptide bonds with respect to each other lead to characteristic CD signals thereby allowing identification of the secondary structural elements.
Materials:
Equipments:
- Circular dichroism spectropolarimeter
- Weighing balance
- pH meter
Reagents:
- 50 mM phosphate buffer, pH 7.0
- Given protein (Hen egg white lysozyme)
- 0.1 M KCl (Note 1)
Glassware and plasticware:
- Pipettes
- Pipette tips
- 100 ml volumetric flasks
- 100 ml beaker
- Test tubes or 15 ml polypropylene tubes
- 1 mm path length Quartz cuvettes
Preparation of reagents:
Phosphate buffer (Note 1) :
- Prepare 50 mM sodium phosphate buffer, pH 7.0 as described in lecture 3.
- Filter the buffer through a 0.22 μm filter.
Protein solution (Note 2):
- Switch on the UV/visible spectrophotometer and allow it 30 minutes warm up.
- Meanwhile, weigh 5 mg lysozyme and dissolve it in 500 μl phosphate buffer.
- Filter the solution through 0.22 micron filter.
- Take 10 μl of the filtered lysozyme solution and add 990 μl of 0.1 M KCl.
- Measure the absorbance of the 100-fold diluted lysozyme solution against 0.1 M KCl at 281.5 nm
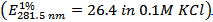 .
.
- Estimate the concentration of the lysozyme stock solution using the formula:
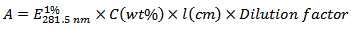
- Dilute the lysozyme stock solution to prepare a 20 μg/ml solution in 50 mM phosphate buffer, pH 7.0. This is the working protein solution.