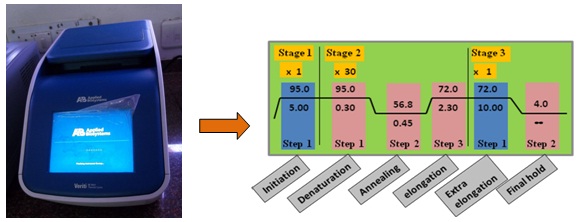
Figure 15.2. Representation of thermal cycler instrument showing the position of sample and schematic diagram of 30 cycle PCR.
Two restriction enzymes sites are added at the 5' end of each of the primer to facilitate cloning. The chosen restriction enzymes will not cut DNA fragment (non-cutters). Typically 3 to 4 nucleotides are added at the end of the restriction sites to allow efficient cutting by restriction enzymes.
Primer Designing and criteria: For a specific amplifications in PCR, good primer design is essential. The following parameters needs to be considered while designing a primer:
1. Primer length: Oligonucleotides between 18-24 bases is the ideal length which is long enough for adequate specificity and short enough for primers to bind easily to the template at the annealing temperature.
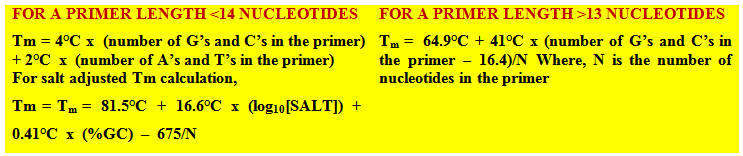
2. Primer melting temperature (Tm): Primers with melting temperatures in the range of 52-58oC generally gives the best results. The GC content of the sequence gives a fair indication of the primer Tm. The two primers should be prepared in such a way that their Tm difference should not be more than 2°C otherwise it will result in poor annealing efficiency. Tm can be calculated by the following formula:
