C. Elongation: This is the synthesis step where the polymerase perform synthesis of new strand in the 5’ to 3’ direction using primer and deoxyribonucleoside triphosphates (dNTPs). An average DNA polymerase adds about 1,000 bp/minute. Step 1,2,3 makes one cycle and in general 35-40 such cycles are performed in a typical PCR amplification.
2. After the cycles are completed, the reaction is held at 70-74°C for several minutes to allow final extension of the remaining DNA to be fully extended.
3. The reaction is complete and the resulting amplified nucleic acids are held at a low temperature (~4°C) until analysis.
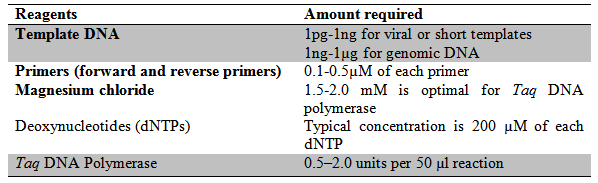
Reagents: The reagents required for a complete PCR reaction volume is given in the table
Instrumentation: Thermal cycler is the instrument that carries out the amplification via polymerase chain reaction (Figure 15.2). Usually the three main events are repeated for 30-40 cycles to obtain detectable amount of product at the end of the cycles. The automated system performs the cyclic temperature changes required for enzymatic amplification of specific DNA segments in vitro using this PCR. The device has a thermal block with holes where tubes containing reaction mixtures can be inserted. The cycler varies the temperature of the block in discrete, pre-programmed steps using peltier effect.
Primers: A primer is a short oligonucleotide that serves as a starting point for DNA synthesis. In PCR, two primers are required to bind to each of the single stranded DNA (obtained after denaturation) flanking the target sequence. These are called Forward and Reverse primers. They primers are designed in such a way that they have a sequence complimentary to the sequence in the template DNA.