CD of nucleic acids
As mentioned in lecture 5, nitrogenous bases constitute the chromophores of nucleic acids in the near and far UV region. The CD of the stacked bases is larger in magnitude as compared to that of the isolated bases. As the double helical nucleic acids have stacked bases, what we measure essentially is the CD that arises due to coupling of the chromophores. As the stacking geometries are different for different forms of nucleic acids such as B-DNA, Z-DNA, and A DNA; CD can help in determining which DNA form is present in a given sample.
Applications in biomolecular analysis
- Determination of protein/peptide structure: As has already been discussed earlier, far UV CD spectroscopy provides information about the secondary structural elements in a protein. A mixture of structures can be deconvoluted to obtain the fraction of different structural elements. Furthermore, near UV CD provides information about the tertiary structure of the protein.
- Comparison of structures: Mutants of proteins are often required for understanding the functions of the proteins. It, however, needs to be ascertained that the mutation does not cause any significant change in the overall structure of the protein. CD spectroscopy happens to be a fast and extremely reliable tool to compare the conformations of the wild type proteins with their mutants.
- Stability of proteins: Stability of the proteins to denaturants or heat can be studied using CD spectroscopy. In such studies CD is usually monitored at a single wavelength, typically around 220 nm. Plotting the change in ellipticity against increasing denaturant concentration/temperature provides the denaturation curve. Figure 9.3 shows the denaturation curves for three related proteins. The denaturation curves suggest that the protein indicated with the blue trace is most stable while the one indicated with red trace the least.
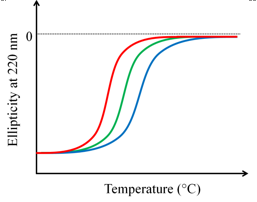
Figure 9.3 Comparison of thermostability of three related proteins. The blue trace represents the most stable protein.