Discovery of polymerase chain reaction (PCR) (Figure 2.2) was a major step forward in the biomedical research and diagnostics. Presence of a pathogen inside the body is classically detected using serological methods or culture of the infectious agents. Owing to its excellent sensitivity, PCR can detect the presence of pathogens earlier than the serological tests. Other than infectious diseases, PCR is also used to detect genetic disorders. It is hard to imagine doing research in the areas of molecular genetics without employing PCR.
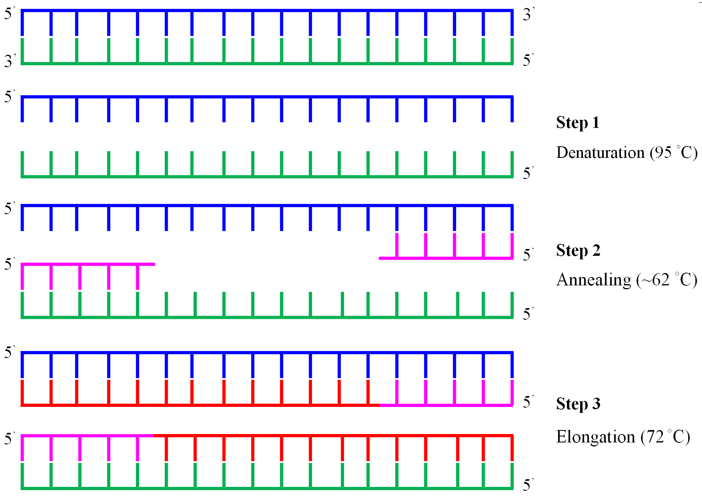
Figure 2.2 Principle of PCR amplification of DNA
Complexity of the biological systems hardly needs any mention. To understand the molecules at function in a living system, it is important to look at the system altogether rather than individual components and processes. Sequencing of complete genomes led researchers to estimate the number of genes a particular organism expresses and further to understand the co-expression of a large number of genes and their role in physiology and pathophysiology. Identification and estimation of the subset of proteins expressed at any instant can provide useful information about the system viz. expression of a gene or a set of genes beyond a threshold level may be a marker of a disease. Intrinsically low levels of a large number of proteins, however, posed a challenge for detecting and identifying them. Application of mass spectrometry to proteins and peptides provided major breakthrough towards achieving this. Mild ionization techniques such as electrospray ionization (ESI) and matrix assisted laser desorption ionization (MALDI) could successfully ionize large biomolecules without damaging them. This opened up a plethora of possibilities and resulted in the development of a new research discipline, called proteomics . Proteomics refers to the study of the complete set of proteins expressed by a cell or an organism. The ultimate goal of the proteomic studies is to identify all the proteins present in the specimen; quantify them; and identify the posttranslational modifications, if any. A proteomic approach typically starts with the isolation of the total protein from the sample. Total protein is then resolved into its components using 2-D gel electrophoresis that separates the proteins according to their isoelectric points in one dimension and their molecular weights in the other. The individual protein spots are then cut from the gel and eluted out. The proteins are then analyzed using mass spectrometry either directly or after digesting with a sequence specific protease such as trypsin. The proteins can then be identified either by de novo sequencing or by using databases having sequence information and thereby mass information of the peptide fragments (discussed in lecture 13). Proteomic analysis is useful in identifying the markers for various processes and diseases. For example, comparing the samples from a set of healthy individuals with that of individuals having some disease/disorder can identify if the protein levels go up or down in the unhealthy individuals as compared to the healthy ones. Systematic studies with a large number of individuals are likely to result in identification of biomarkers for the diseases. The need to retrieve and analyze the huge amount of data generated from genome sequencing projects led to the development of another discipline, called Bioinformatics. Bioinformatics utilizes computer science and mathematics to organize and retrieve the biological data. The biological information such as sequences of nucleic acids and proteins, their structures, post-translational modifications of proteins, etc. are organized and stored in the databases. The databases can be accessed to retrieve the required information for analysis.