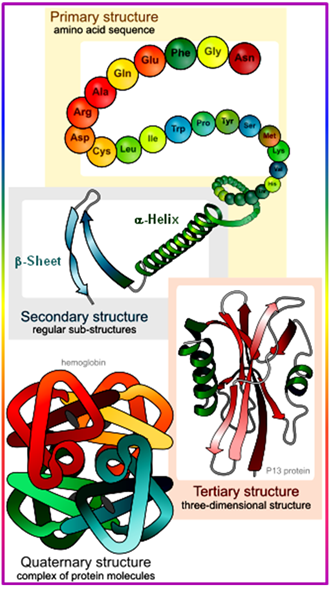
2.4. Peptide Secondary Structures and Tools for Stabilization
ΦCα — NΨCα — C |
|
Thus we can calculate the backbone conformations of a peptide through interplay of rotation around the bonds defined by the torsion angles Phi (Φ) and Psi (Ψ) and the steric hindrance of side groups determined by their Van der Waals radii. The resulting conformational map is called Ramachandran plot. A free rotation because of thermal motion around a C-C bond is possible in the absence of any steric i.e., Van der Waals radii constraints.
- Tertiary (3°) Structure: three-dimensional arrangement of all atoms in a single polypeptide chain is the main feature of tertiary structure. The tertiary structure is the 3-dimensional, native structure of a single polypeptide or protein. A protein normally is folded into a compact structure. The secondary structures are stabilized by the final, native fold. The native fold is defined as the active conformation which is mostly stabilized by H-bonding, Electrostatic or Ionic, Hydrophobic and Van der Waals forces of interaction and some time by Covalent interaction.
The location of an amino acid in a protein fold correlates with the energy of hydration of each individual amino acid residue and the entropy of the side chain, polypeptide backbone, and solvent molecules. As mentioned earlier, polar and charged amino acids are likely to be hydrated, whereas the non-polar residues stick to each other and often form the core of a protein, forming a usually hydrophobic core that stabilizes the fold of water soluble, globular proteins. For membrane proteins exhibiting both hydrophobic and hydrophilic surfaces, the amino acid distribution is different from globular proteins but the same rules of the hydrophobic effect apply.
- Quaternary (4°) structure: It is the overall organization of non-covalently linked subunits of a functional protein. Single polypeptides can associate with each other to form larger protein complexes of geometrically specific arrangements, called quaternary structures. Individual polypeptides in protein complexes are referred to as subunits. Most enzymes are complexes of proteins and the symmetry and stoichiometry of the composition of the complexes is crucial for their activity.
One can distinguish two different compositions, the homomeric and heteromeric complexes. Heteromeric composition of most protein complexes gives the cells an additional level of variability and complexity it can use for its activity. Often, heteromeric compositions of protein complexes are tissue specific or developmental specific and multiple genes can control the activity of a single heteromeric protein complex.
- Primary Structural Motifs:
- α-Helix: a right handed helical structure with average torsion angles Φ = -57 and Y = -47
- β-Sheet: parallel (Φ = -119 and Y = 113) or anti-parallel pleated sheet structures
- β-Turn: minimal loop structures of 3 to 4 amino acids with defined torsion angles
- Disulfide bonds: Peptide or proteins may form covalent linkage between two cysteine amino acids.
- Forces Stabilizing Peptide Structures:
- H-bonding
- Electrostatic or Ionic
- Hydrophobic and Van der Waals
- Covalent