Screening of Recombinant Clone
Lab Experiment 40.3 : Verification of recombinant DNA (Colony PCR)
Principle of the technique: The whole process of PCR involves three main events, Denaturation, Annealing and Elongation (Figure 40.1). A DNA fragment of interest is used as a template and a pair of primers which are short oligonucleotides complimentary to the both strands of the template DNA. The purpose of primer is to initiate the DNA synthesis in the direction of 5’ to 3’. The number of amplified DNA or the amplicons increases exponentially per cycle thus one molecule of DNA give rise to 2,4,8,16 and so forth. This continuous doubling is carried out by a specific enzyme called DNA polymerase which sits at the unfinished double stranded DNA created by template DNA and primer. For further extension of the DNA, the polymerase enzyme require supply of other DNA-building blocks such as the nucleotides consisting of four bases Adenine (A), Thymine (T), Cytosine (C) and Guanine (G). The template, primer, polymerase and four bases are the main components for polymerase chain reaction.
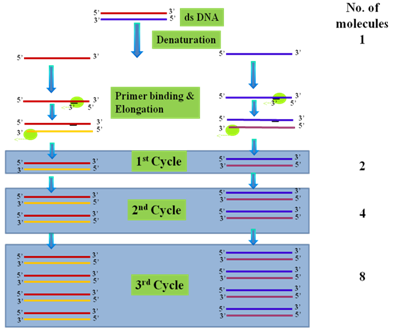
Figure 40.1: Basic Principle of polymerase chain reaction (PCR).
Instrumentation: Thermal cycler is the instrument that carries out the amplification via polymerase chain reaction (Figure 40.2). Usually the three main events are repeated for 30-40 cycles to obtain detectable amount of product at the end of the cycles. The automated system performs the cyclic temperature changes required for enzymatic amplification of specific DNA segments in vitro using this PCR. The device has a thermal block with holes where tubes containing reaction mixtures can be inserted. The cycler varies the temperature of the block in discrete, pre-programmed steps using peltier effect.
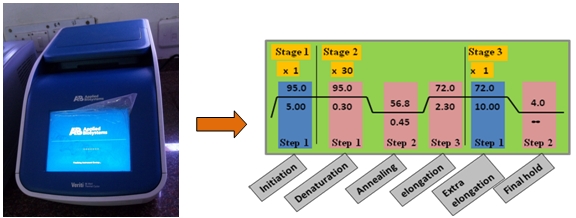
Figure 40.2. Representation of thermal cycler instrument showing the position of sample and schematic diagram of 30 cycle PCR.