Transcription activation & repression
- There are multiple modes of transcription activation. All these usually involve different protein factors (activators) that bind DNA sequences (enhancer sequences) in and around the promoter site. All these act to increase the affinity of the initiation complex or RNA holoenzyme at the promoter (thus enhancing the chance that a productive open complex will form and transcription will initiate).
- When enhancer elements combine with poor promoter sequences, activators can modulate the activity of a gene by several hundred or thousand fold. For example, if a gene has a poor -35 or -10 box in its promoter site, it will be poorly expressed - an activator protein can, in such a case, enhance the activity of the gene by several orders of magnitude, by recruiting the RNA polymerase (or other initiation factors) to the promoter site.
- Repression, on the other hand, works by having protein (repressors) that sit on or close to the promoter regions of the DNA, preventing RNA polymerase or initiator/activator proteins from starting transcription initiation (thereby "turning-off" the gene).
- Activators and repressors are usually DNA binding proteins. These proteins have common DNA binding motifs such as Zn-fingers, helix-turn-helix, etc. Activators also have regions that interact with different domains of the RNA polymerase (parts of alpha, sigma, etc.).
- In addition to protein activators, there are also DNA sequences that can directly interact with RNA polymerase components. For example, the alpha subunit in the bacterial polymerase has two domains - the alpha NTD (n-terminal domain) and the alpha CTD (c-terminal domain), linked by a flexible linker region. The alpha CTD can bind DNA sequences (UP elements) upstream of the promoter region, enhancing the affinity of the polymerase on the promoter.
Transcription termination
- Termination in bacterial system can be broadly classified as "rho independent", and "rho dependent".
- In rho independent termination, there is formation of a stable GC rich stem-loop in the newly synthesized RNA followed by a string of U's (A's in the template strand) spaced about 20 bases downstream (these sites are often called intrinsic terminators). The stem loop "snares" the polymerase, slowing or stalling it. This pause, coupled with the low stability of the RNA-DNA hybrid at the active site (run of A=U basepairs) allows the RNA polymerase to fall off the template DNA and terminates the RNA transcription for that gene.
- In rho dependent termination, rho binds the newly formed RNA (as hexamers), and stalls the RNA polymerase by interacting with it. In some models, the Rho hexamer translocates on the RNA. Rho termination activity is stimulated by ATP hydrolysis. This activity is greatly enhanced by Nus factors.
- In eukaryotic cells, transcription termination involves cleavage of the elongating RNA chain by specific endonucleases which recognize particular sequences (AAUAAA) in the newly formed RNA. Once this happens, the RNA elongation complex is destabilized, and falls off the DNA. It is then available to attach to another nearby PIC, and start transcribing again.
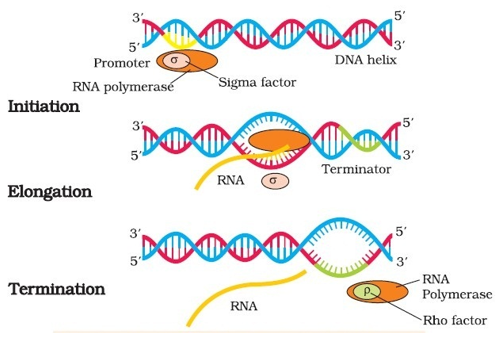
Fig. 13. Process of transcription in bacteria