Transcription initiation
- Transcription initiation in bacteria (prokaryotes) involves sigma factors. The sigma factor combine with the core RNA polymerase to form a holoenzyme that is competent for promoter binding. The core RNA polymerase, by itself, cannot bind the promoter site.
- The sigma factor can be thought of as the specificity factor in the RNA polymerase. Each bacteria has several different sigma factors that recognize slightly different promoter sequences. Predominant among these is the sigma70 (70 kDa in E. coli; also called rpoD), which initiates the transcription of most genes in exponentially growing cells. There are two general classes of sigma factors - sigma 70 class and sigma 54 class:
- The sigma 70 class of sigma factors share extensive sequence homology, and bind to two conserved sequences upstream of the gene start site (the -35 box and the -10 box). Each of these sigma factors recognizes slight variations in these conserved sequence boxes. Sigma70 binds promoter DNA as a part of the holoenzyme, and binds DNA very poorly in absence of the core enzyme.
- The sigma54 class of genes (54 kDa in E. coli; also called rpoN) controls a much smaller set of genes than sigma70. It recognizes different conserved sequences (the -12 and -24 boxes). Unlike Sigma70, sigma54 can bind DNA even in the absence of the core RNA polymerase. It however lacks the ability to melt promoter DNA on its own - for this it needs to interact with other activator proteins that bind further upstream of the promoter site, as well as the core RNAP.
- In eukaryotic systems, transcription initiation is very different. There are no sigma factors. Instead, the central protein in forming the "pre-initiation complex " (PIC) is the TATA binding protein (TBP), that binds to the TATA box, a conserved sequence just upstream of the initiation region. A large number of other general transcription factors such as TFIIB, TFIIE, TFIIF, TFIIH (TF stands for transcription factor; II stands for Pol II; there are similar factors for Pol I and Pol II) and others assemble to form the multisubunit TFIID complex. This PIC then recruits the RNA polymerase (Pol I, II, or III in eukaryotic cells) to initiate transcription. The PIC often remains at the promoter site, and is then available to initiate another round of transcription.
- TBP is a universal transcription factor, and is seen in all eukaryotes and archaea. It sharply bends DNA at the TATA box.
Transcription elongation
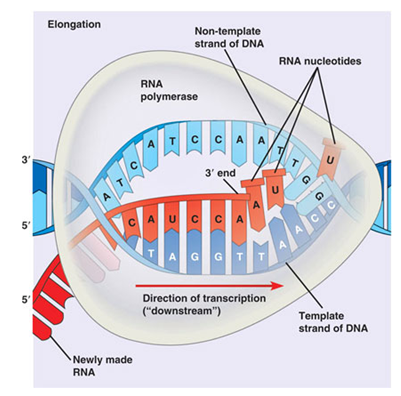
- Once initiation is complete and the open complex forms, the RNA polymerase begins to read the template strand and add corresponding RNA nucleotides. This process is not always efficient, and the polymerase may make several passes at this. After a long enough RNA-DNA hybrid is made, the polymerase clears the promoter region and moves rapidly downstream. This is preceeded by a large conformational change in the polymerase core enzyme, as it clamps down on the DNA and becomes quite processive.
- While transcription elongation is quite rapid, the polymerase does not transcribe all sequences with equal efficieny. The elongation rate is not uniform, and there can be pausing or stalling. Elongation factors (GreA/GreB in bacterial systems; TFIIS in eukaryotes) act to help the polymerase along by stimulating backtracking and cleavage of the newly formed RNA (from the 3' end).
- Other factors are involved in the elongation cycle - for example, in eukaryotic Pol II there are the elongins and ELL proteins that increase the elongation rate, as well as factors to remodel chromatin.
 ]
]
Fig. 12. Transcription elongation