Ribosomal Proteins:
A ribosome is composed of three (in bacteria) or four (in eukaryotes) different rRNA molecules and as many as 83 proteins, organized into a large subunit and a small subunit. The primary structure of several of these proteins has been elucidated. Most of the recent knowledge about the structure of ribosomal proteins has been achieved by dissociation of ribosomal subunits into their component rRNA and protein molecules. When both 50S and 30S ribosomal subunits are dissociated by centrifuging both of them in a gradient of 5 M cesium chloride, then there are two inactive core particles (40S and 23S, respectively) which contain the RNA and some proteins called core proteins (CP) at the same time several other proteins—the so-called split proteins (SP) are released from each particle (Fig. 14.3). There are SP50 and SP30 proteins which may reconstitute the functional ribosomal subunit when added to their corresponding core. Some of the split proteins are apparently specific for each ribosomal subunit. The split proteins have been further fractionated and divided into acidic (A) and basic (B) proteins. According to Nomura (1968, 1973) and Garett and Wittmann (1973) each 70S ribosome of E. coli is composed of about 55 ribosomal proteins. Out of these 55 proteins, about 21 different molecules have been isolated from the 30S ribosomal subunit, and some 32 to 34 proteins from the 50S ribosomal subunit. Similar organization of ribosomal proteins and RNA is found in 80S Ribosomes. Different rRNA molecules evidently play a central role in the catalytic activities of ribosomes in the process of protein synthesis.
Metallic Ions:
The most important low molecular weight components of ribosomes are the divalent metallic ions such as Mg++, Ca++ and Mn++.
Structure
The ribosomes are oblate spheroid structures of 150 to 250 Ao in diameter. Each ribosome is porous, hydrated and composed of two subunits. One ribosomal subunit is large in size and has a domelike shape, while the other ribosomal subunit is smaller in size, occurring above the larger subunit and forming a cap-like structure. The small ribosomal subunit contains a single rRNA molecule, referred to as small rRNA. The large subunit contains a molecule of large rRNA and one molecule of 5S rRNA, plus an additional molecule of 5.8S rRNA in vertebrates. The lengths of the rRNA molecules, the quantity of proteins in each subunit, and consequently the sizes of the subunits differ in bacterial and eukaryotic cells. The assembled ribosome is 70S in bacteria and 80S in vertebrates. There are great structural and functional similarities between ribosomes from all species which is another reflection of the common evolutionary origin of the most basic constituents of living cells.
The 70S ribosome consists of two subunits, 50S and 30S. The 50S ribosomal subunit is larger in size and has the size of 160 Ao to 180 Ao. The 30S ribosomal subunit is smaller in size and occurs above the 50S subunit like a cap. The 80S ribosome also consists of two subunits, 60S and 40S. The 60S ribosomal subunit is dome-shaped and larger in size. In the ribosomes which remain attached with the membranes of endoplasmic reticulum and nucleus, the 60S subunit remains attached with the membranes. The 40S ribosomal subunit is smaller in size and occurs above the 60s subunit forming a cap-like structure. Both the subunits remain separated by a narrow cleft. The two ribosomal subunits remain united with each other due to high concentration of the Mg++(.001M) ions. When the concentration of Mg++ions reduces in the matrix, both ribosomal subunits get separated. Actually in bacterial cells the two subunits are found to occur freely in the cytoplasm and they unite only during the process of protein synthesis. At high concentration of Mg++ ions in the
matrix, the two ribosomes (monosomes) become associated with each other and known as the
dimer. Further, during protein synthesis many ribosomes are aggregated due to common messenger RNA and form the polyribosomes or polysomes.
The actual three-dimensional structures of bacterial rRNAs from Thermus thermopolis recently have been determined by x-ray crystallography of the 70S ribosome. The multiple, much smaller ribosomal proteins for the most part are associated with the surface of the rRNAs. During translation, a ribosome moves along an mRNA chain, interacting with various protein factors and tRNAs and very likely undergoing large conformational changes (see Figure 2). Despite the complexity of the ribosome, great progress has been made in determining the overall structure of bacterial ribosomes and in identifying various reactive sites. X-ray crystallographic studies on the T. thermophilus 70S ribosome, for instance, not only have revealed the dimensions and overall shape of the ribosomal subunits but also have localized the positions of tRNAs bound to the ribosome during elongation of a growing protein chain. In addition, powerful chemical techniques such as footprinting, have been used to identify specific nucleotide sequences in rRNAs that bind to protein or another RNA. Figure 1 illustrates the ribosomes.
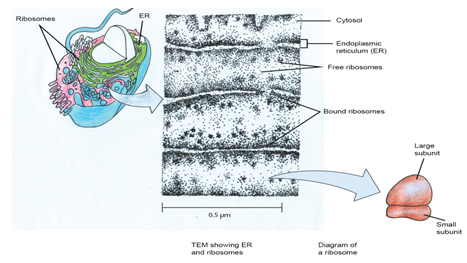
Figure 1: Schematic representation of the ribosome.