3. DNA gyrase: This enzyme is required to unwind the double stranded DNA.
4. DNA ligase: The enzyme is required to join the DNA fragments.
5. Single Stranded DNA Binding (SSB) Protein: After unwind DNA, SSB binds to the single stranded DNA to avoid rewinding. It helps to reduce the energy required for unwinding.
In addition, replication machinery needs to complete this task in 4 steps:
1. Identify the site of occurrence: it is difficult to start the replication of circular or very large DNA fragment randomly. Hence, a particular pattern of nucleotide sequence exists on DNA to recognize and facilitates the replication initiation. The difficult task with regard to replication is to denature the double stranded DNA and presence of sequence with low melting temperature will facilitate the event. As discussed in earlier lecture, interaction between adenine and thymine is mediated by 2 hydrogen bonding whereas guanine and cytosine is mediated by three hydrogen bonding. As a result presence of AT rich region will facilitate earlier denaturation and assembly of replication machinery. These sequences are mostly present in a region and known as origin of replication. A typical example of E.coli origin of replication is given in the Figure 27.1.
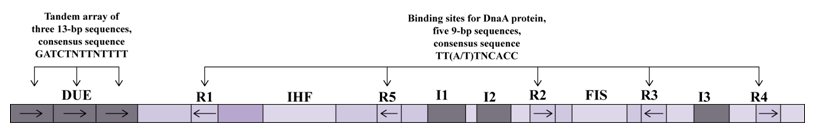
Figure 27.1: E.Coli Origin of Replication.
2. Initiation: The unzipping of DNA at the origin of replication forms a “Y” shaped structure known as replication fork (Figure 27.2). A short chain of RNA is formed at the 5´end. This is a RNA primer and synthesized by primase. Synthesis of RNA primer is essential as DNA polymerase cannot perform denovo DNA synthesis.
3. Elongation: In the elongation step, DNA polymerase start the synthesis of DNA on the pre-existing short oligomeric nucleotide strand. In this step, an incoming deoxynucleotide triphosphate get joined by hydrogen bonding to the appropriate nitrogen bases of the single DNA chain as per the base pairing rule; A-T, T-A, C-G and G-C. The nucleotide triphosphate joined to each DNA strand, break off their high energy bonds and set free pyrophosphate (P-P) molecules. Pyrophosphate undergoes hydrolysis with the help of an enzyme pyrophosphatase, and release energy and set free inorganic phosphate group 2Pi. The energy released in this process is used to derive the polymerization of nucleoside to form DNA (Eq 27.1). The released deoxynucleotide monophosphate joined to each single DNA chain become linked together to form new DNA chain. DNA polymerase.
| 27.1 |