2.8.2. Methodology for Genetic incorporation of UNAA:
The historical development of synthesis or semisynthetic methods for introduction of unnatural amino acids into peptide and proteins[(a) Offord, R. E. (1987) Protein Eng. 1,151-157; (b) Kaiser, E. T. (1988) Angew. Chem. Int. Ed.Engl. 27,913-922] by Offord and Kaiser has paved the way to develop methods for site specific incorporation of more unnatural amino acids into proteins by Schultz et al. [Corey, D. R. and Schultz, P. G. (1987) Science 238, 1401-1403] towards expanding the genetic code and thereby to give rise to the birth of semisynthetic organism. Towards this goal, an increasing amount of interest from various research groups has resulted in the acceleration of progress of design of unnatural amino acids for application in protein engineering. Several non natural amino acids were reported looking after the steric and electronic properties and incorporated into proteins site specifically. However, many of the reported unnatural amino acids are not suitable for giving novel biological properties of the proteins or not containing functionality for labeling the proteins. Moreover, ffluorescently labeled proteins are useful in a large number of bioanalytical applications, but, a little was attempted to develop fluorescent unnatural amino acids (FUAA) or fluorescently labelled UAA for genetic encoding or to generate labelled proteins/peptide for studying conformational or diverse functional realm.
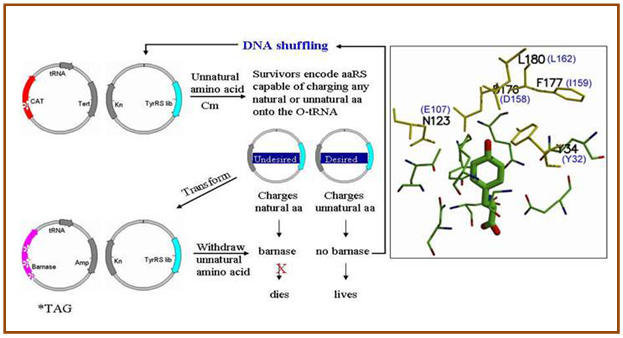
Figure 2.60: Scheme for evolving aminoacyl tRNA synthetases with novel specificities.
Towards this end, methodologies were developed that allow one to genetically encode novel amino acids, beyond the common twenty, in prokaryotic and eukaryotic organisms. One of such methodologies involves the generation a unique codon-tRNA pair and corresponding aminoacyl-tRNA synthetase. Specifically, an orthogonal tRNA is constructed that is not a substrate for any natural aminoacyl synthetases and which inserts its cognate amino acid in response to the amber nonsense codon. A cognate synthetase is then generated which recognizes this unique tRNA and no other; the substrate specificity of this synthetase is then evolved to recognize a desired “twenty first” amino acid, and no endogenous amino acid. Sultz et al. have shown that this methodology can be used to efficiently incorporate a large number of amino acids into proteins in E. coli and yeast with fidelity and efficiency rivaling that of the common amino acids. Using this methodology, several research groups have added a variety of novel amino acids to the genetic codes of E. coli.
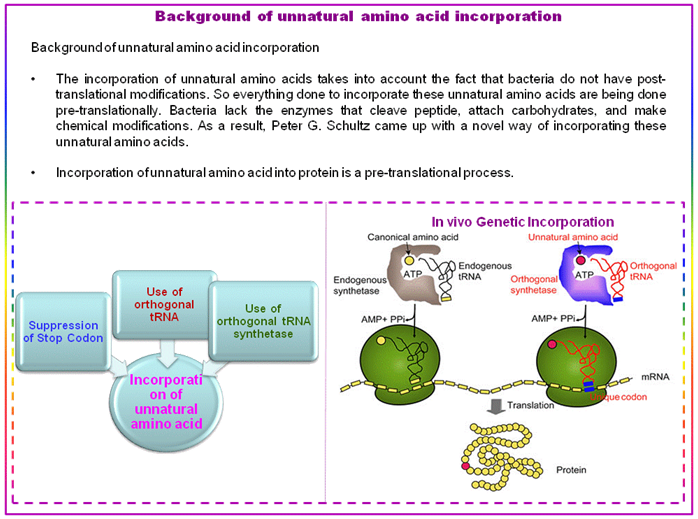
Figure 2.61: Scheme for in vivo incorporation of unnatural amino acids.