2.8. Expanding the Genetic Code
2.8.1. Background:
The tools of chemistry, most notably chemical synthesis and spectroscopy, have had a remarkable impact on biology-from the structural elucidation of the double helix to the chemical synthesis of peptides and oligonucleotides. At the same time, modern molecular biology has made it possible not only to manipulate protein and nucleic acid structure but also the genetic composition of living organisms. The ability to use these tools in combination opens an unprecedented opportunity in the coming millennium, both for understanding complex biological systems at a molecular level as well as for the generation of molecules with novel biological, chemical and physical properties.
Proteins play a vital role in all living organisms to maintain the cell structures, properties and functions which is again dependent on post translational modifications. The genetic codes of every known organism encode the same 20 amino acid building blocks using triplet codons generated from A, G, C and T. Thus, in all organisms, the building blocks of all the translated proteins are the same 20 natural amino acids. Natural selection/evolution has generated a large no of proteins with more or less common structures and functions in a population. Therefore, these proteins are highly specialized for specific functions and thus, are not suitable for a different function other than they use to do the specific job. To perform a complex additional function by a protein, it needs other functionalities within its framework. Posttranslational modifications, cofactor-dependent catalysis, and pyrrolysine/selenocysteine incorporation in bacteria proves that the natural evolutionary movement needs extra chemical functionalities other than those present within the 20 natural amino acids. Then the knowledge from the nature knocked scientists to ask oneself: Why only this set of amino acids and not any additional ones are used for genetic coding? Is this the ideal number? How did they determine the complex folds and functions of proteins? Would additional amino acids allow the generation of proteins or even entire organisms with enhanced properties? These questions need yet to be fully answered.
The ability to generate proteins with new building blocks, beyond those specified by the genetic code, would not only provide a powerful tool with which to investigate these questions but might also allow us to generate proteins or even entire organisms with novel functions. The ability to introduce amino acids with precisely tailored steric and electronic properties into proteins would also allow us to carry out “physical organic” studies of proteins much the same way as has been historically done with small molecules.
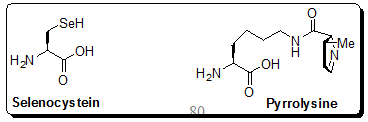
Figure 2.58: Presentation of naturally occurring 21st and 22nd amino acids
Thus, the approach, “directed evolution” have come up with proteins of altered structural and functional properties that do not occur in nature. As for example, researchers are trying to generate new proteins to function as potent therapeutic, microbial enzymes for fuel production from agricultural waste, imaging and ultimately semi-synthetic organism with diverse functionality. Now a day, scientists are involved in designing the proteins via rational approach of directed evolution to produce new proteins with desirable properties. Therefore, inspired by Natures’ post-translational modification to include different functional units or molecules into the proteins, chemoselective conjugation methods have been developed to attach probe to proteins; thereby facilitating the study of structure and functions at molecular level. However, conventional bioconjugation reaction has several drawbacks and mostly exploited the only nine canonical aminoacids with limited functional groups and abundance for modification/ligation within a protein. Therefore, because of that, site-specific conjugation to a desired canonical amino acid in a protein is difficult task. To circumvent this problem, several chemical and biochemical methodology have been developed to site-specifically incorporate designer amino acids (the unnatural amino acids) with disered functionalities to probe protein structure and function and to generate proteins/enzyme for several novel biochemical applications, such as for chemical synthesis, biomedical research or even as therapeutics. Therefore, the research in the field of design and synthesis of non-natural amino acids with novel properties, to encode it genetically and to incorporate it site-specifically into a protein via bio-orthogonal strategy, is growing at a fast space for the growing demand of proteins of potential therapeutic and many other diversified novel functional applications. Therefore, an expanded genetic code, site-specifically incorporated into protein would allow us to study proteins’ physical organics which would otherwise be extremely difficult-(a) probing protein structure, function and interaction, (b) regulating protein activity, (c) monitoring the mode of action, (d) improving immunogenicity, and (e) very recent development of a protein with a “chemical warhead” which target specific cellular components.
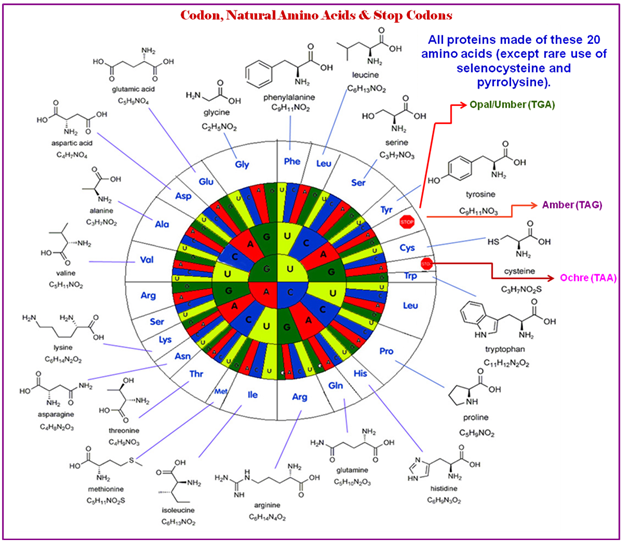
Figure 2.59: The structures of the amino acids and their codons.