1 . Restriction enzyme
Restriction enzymes are enzymes isolated from bacteria that recognize specific sequences in DNA and then cut the DNA to produce fragments, called restriction fragments. Because they cut within the molecule, they are often called restriction endonucleases. Detection of sequences varies between 4 and 8 nucleotides, it may be of palindromic, which correspond to nitrogenous base sequences that read the same backward and forward.
Examples
a) EcoR1 restriction cleavage produces "sticky" ends,
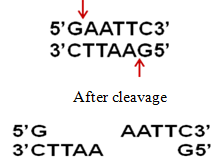
b) Sma I restriction enzyme cleavage produces "blunt" ends:
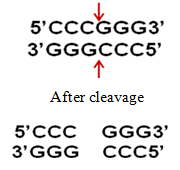
Restriction enzymes play a very important role in the construction of recombinant DNA molecules, as is done in gene cloning.
2. Types of restriction enzyme
Restriction enzymes are classified into four general groups (Types I, II, III and IV) based on their composition and requirements of enzyme cofactor, the nature of their target sequence, and the position of their DNA cleavage site relative to the target sequence:
1. Type I enzymes
Cleave at sites remote from recognition site. Both ATP and S-adenosyl-L-methionine is required for cleavage. These enzymes are multifunctional and are capable of both restriction and modification activities, depending upon the methylation status of the target DNA.
2. Type II enzymes
Cleave within or at short specific distances from recognition site. It requires mostly magnesium as cofactor. The enzyme perform single function that is restriction. This type of enzymes is independent of methylase.
3. Type III enzymes
Cleave at sites a short distance from recognition site. It requires ATP (but doesn't hydrolyse it); S-adenosyl-L-methionine stimulates reaction but is not required. It exists as part of a complex with a modification methylase
4. Type IV enzymes targets on methylated DNA.
Some of the common IV restriction enzymes are R. Eco 57I, R. Bce 83I, R. Hae IV, R. Mme I, R. Bsp LU11III and Bse MII. The common feature of these enzymes is that two enzymatic activities, methyltransferase (MTase) and endonuclease (ENase), are combined together in one polypeptide chain and the ENase activity is positively affected by S -adenosine- l -methionine (AdoMet) but ATP has no influence on activity of the enzymes. As for their recognition pattern, type IV enzymes usually recognize asymmetrical DNA sequences and ENase cleavage occurs at a defined distance from the recognition site, as in type IIS enzymes.