Primer:
In order to perform the sequencing, the first step is the annealing of a gene specific or a universal oligonucleotide primer to the recombinant M13 vector or single stranded DNA. This primer acts as a starting point for the DNA polymerases enzymes for the complementary strand synthesis. Primer is generally radiolabelled to generate the autoradiograph. The label may also introduce into the new strand by adding radiolabelled deoxynucleotides e.g. S 35 or P32 in the reaction mixture (Fig.2).
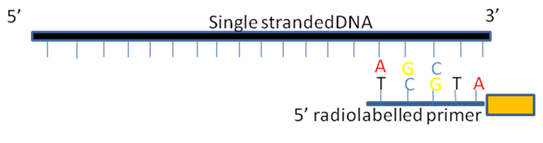
Figure 2. Annealing of primer to the Template DNA.
Enzyme:
To generate a DNA ladder, enzymes are being used that lacks 3'-5' exonuclease activity. Previously Klenow fragment was in use for this purpose and DNA synthesis was done at 37 0 C which was the optimum temperature for DNA polymerase I. DNA polymerase adds either deoxynucleotide or the corresponding 2',3' dideoxynucleotide form at the extension step. Presently Sequenase, a modified bacteriophage T7 DNA polymerase has almost replaced the Klenow fragment. Sequenase has higher processivity and it generates bands with similar intensity and a readable DNA sequence while Klenow has a lower processivity only upto 250 bp. Sequencing polymerases other than sequenases catalyze the incorporation of ddNTPs at only 0.02-1% of the rate of dNTPs but sequenase catalyses the incorporation of ddNTPs at 33% efficiency against its corresponding dNTP. The modified DNA polymerases are formulated in such a way so that they can pass easily through GC stretches and other difficult sequences easily and provide uniform peak heights.
|