DNA Sequence Analysis Methods-I
Download this lecture
Introduction:
Before 1970's there was no direct method to determine the nucleotide sequence. In the mid of 1970's, two methods developed for the direct sequencing of DNA. These were the Sanger Coulson's chain termination method and Maxam Gilbert's chain termination method. For which they shared Nobel Prize in Chemistry (1980).
Sanger method:
Sanger et al. (1974) used the principle of DNA replication for the development of Dideoxy Sequencing method. For the coupling of nucleotides, the 3' hydroxyl group is needed. Sanger used this site to develop the chain termination reaction. He used dideoxyribose in which the hydroxyl group was missing at both 2' and 3' Carbon places in the ribose sugar. These molecules terminate DNA chain elongation because they cannot form a phosphodiester bond with the next deoxynucleotide and the chain proliferation reaction irreversibly stops (Fig. 1).
The chain termination reaction depends on the synthesis of the second strand of the DNA so the template is needed in the single stranded form. Previously the template molecule is generally cloned into M13 vector to get a single stranded DNA and the second strand is enzymatically synthesized. Denaturation with alkali as NaOH, are also being applied to get single stranded DNA.
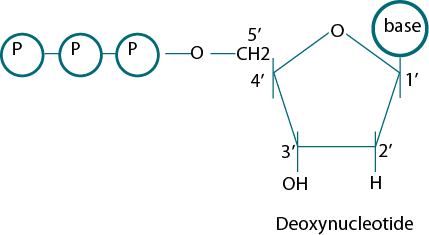
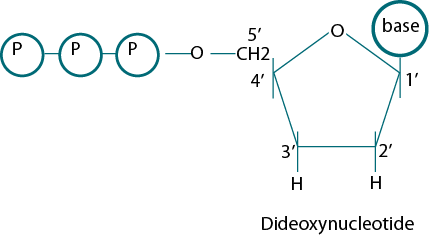
Figure 1: The structure of five carbon sugars deoxy and dideoxynucleotide.
|