A large library of mutants for a given protein gene is made. In these cases, mutagenesis is frequently done using error-prone PCR. This incorporates random mutation in the gene. The peptide display libraries are cloned directly into the viral genome so that all five copies of gpIII display the peptide fusion. Phase displaying the desired sequence is finally selected (Fig. 4).
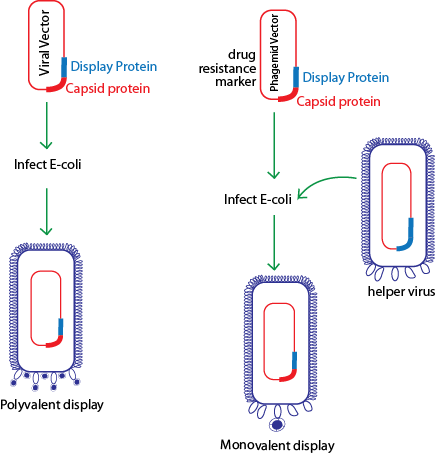
Figure 3: Proteins fused to capsid proteins can be displayed in either of the two formats. Protein can be cloned directly into a viral vector as a fusion to a capsid protein resulting in every copy of the capsid protein displaying the fusion (polyvalency). In case of polyvalent display all gpIII are conjugated with peptide and this results in occasional lost of infectivity. Alternatively, the protein fusion can be constructed in a phagemid vector that carries a copy of the viral capsid gene. In this case, helper phages are mixed at the time of infection. As caspid protein also comes from helper phage (during viral assembly in E.coli ), infectivity of phage displaying peptide is not compromised. However, valency of display of peptide is decreased. This may have some disadvantages at the time of selection. The valancy of display depends on ratio of phage containing phagemid vector and helper phage at the time of infection. Since the helper phage do not carry a drug resistance marker, they are lost upon subsequent selection for drug resistance.
|