1) Proteolysis
During translation, the signal peptide is synthesized first, which is a 20 hydrophobic residue long peptide at the N terminal end of the protein. Once synthesized, it is recognized by the signal recognition particle (SRP) and translation is temporarily stopped. The signal peptide-SRP complex binds to SPR receptor on the ER membrane. This results in opening of a trans-membrane pore (translocon). Now translation resumes and peptide is directed into the ER. As the newly synthesized protein is released in the lumen of the ER, signal peptidases cleave peptide sequence. This explains why despite having AUG as start codon which codes for methionine, all the proteins do not have methionine as the N terminal amino acid. Apart from signal peptide, some polypeptide sequence of the protein is also cleaved resulting in the final sequence. For example, in case of insulin it is released in the ER lumen as preproinsulin. After removal of signal sequence it is called proinsulin, which is a further cleaved form residue 31-60 (C Chain) to form mature insulin (Fig. 2).
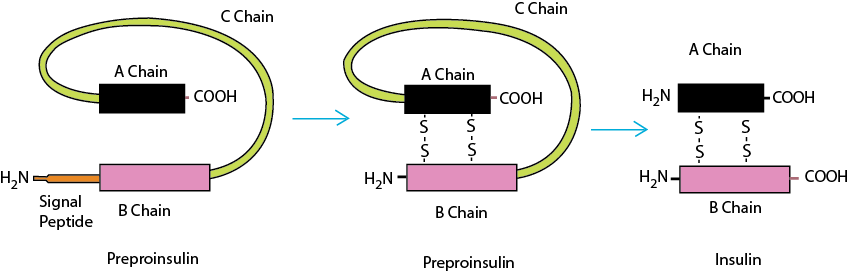
Figure 2: Activation of insulin by proteolysis
2) Glycosylation
Glycosylation is addition of carbohydrate moieties to the polypeptide sequence. Mainly there are two types of glycosylation (Fig. 3)
i) N-linked glycosylation
ii) O-linked glycosylation.
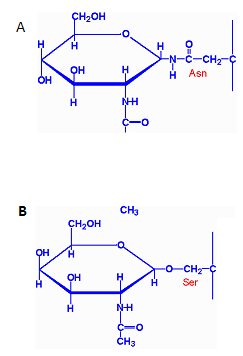
Figure 3: Two types of glycosylation (A) N-linked glycosylation (B) O-linked glycosylation
|