6-5.3(b).3 Sequence Tagged Site (STS) Mapping
Both restriction mapping and FISH face one or more disadvantages. Restriction mapping is easy and rapid technique providing detailed information, but it is not applicable for large genomes. Although FISH can be applied to larger genomes, and fiber-FISH can provide high-resolution data, it is a technically challenging method to implement. STS can overcome many of these limitations and give a detailed map with comparative ease.
STS is an easily recognizable short DNA sequence, typically 100-500 bp in length, generated by PCR using primers obtained from already known sequences. STS (sequence tagged site) markers are used for detailed mapping of large genomes. This method involves collection of various overlapping DNA fragments obtained from either single chromosome or complete genome. The distance between the markers on these fragments determines which fragment contains which STSs (Figure 6-5.3(b).3.1). Closer the markers in the genome, greater will be the chance of occurrence of two STSs on the same fragment.
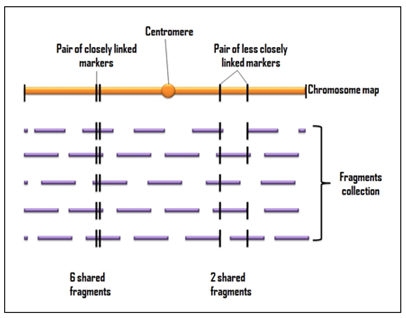
Figure 6-5.3(b).3.1: STS mapping representing the collection of various fragments that span the entire length of a chromosome, with each point on the chromosome present in an average of five fragments. (Adapted and modified form Brown TA. 2002. Genomes. 2 nd ed. Oxford: Wiley-Liss)