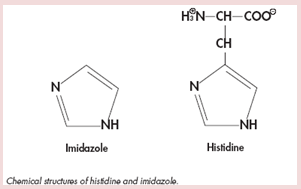
Additional points: As low affinity binding with other contaminant protein is also possible, low concentrations of imidazole in the lysis and wash buffers (10−20 mM) are recommended. The imidazole ring is part of the structure of histidine (Figure shown below). The imidazole rings in the histidine residues of the 6xHis tag bind to the nickel ions immobilized by the NTA groups on the matrix. Imidazole itself can also bind to the nickel ions and disrupt the binding of dispersed histidine residues in nontagged background proteins. At low imidazole concentrations, nonspecific, low affinity binding of background proteins is prevented, while 6xHis-tagged proteins still bind strongly to the Ni-NTA matrix. Therefore, adding imidazole to the lysis buffer leads to greater purity in fewer steps. For most proteins, up to 20 mM imidazole can be used without affecting the yield. If the tagged protein does not bind under these conditions, the amount of imidazole should be reduced to 1−5 mM.
Binding of tagged proteins to Ni-NTA resin is not conformation-dependent and is not affected by most detergents and denaturants. The stability of the 6xHis−Ni-NTA interaction in the presence of low levels of β -ME (up to 20 mM) in the lysis buffer can be used to prevent the copurification of host proteins that may have formed disulfide bonds with the protein of interest during cell lysis. Detergents such as Triton X-100 and Tween 20 (up to 2%), or high salt concentrations (up to 2 M NaCl), also have no effect on binding, and may reduce nonspecific binding to the matrix due to nonspecific hydrophobic or ionic interactions. Nucleic acids that might associate with certain DNA and RNA-binding proteins are also removed without effecting the recovery of the 6xHis-tagged protein .