Use of Nanotechnology : SNPs can also be detected using thin film biosensors (Figure below). An optimized method is described as following:
The biosensors optical layer was prepared by coating the surface of silicon wafers with silicon nitride (Si3N4). This optical layer was also coated with a layer of T structure aminoalkylpolydimethylsiloxane (TSPS) and poly(Phe-Lys) to facilitate covalent attachment of biomolecules. For detection of SNPs a pair of allele-discriminating oligonucleotide probes (blue) and a single detector oligonucleotide (red) is designed. The allele descriminating oligos have 5'-aldehyde groups, a "spacer," followed by 40 nucleotides complementary to the corresponding SNP target sequence. There is difference only at 3'-terminal nucleotide in these probes and this 3'-terminal nucleotide matches with one of the two SNP genotypes. The detector probe has biotin at its 3' end for detection and phosphate at its 5' end for ligation. The assay involves ligation and hybridization of allele discriminating oligomers attached on the biosensor surface followed by detection by antibiotine-antibody. Briefly, target DNA hybridization and ligation of allele descriminationg oligonucleotides and biotinylated detector oligonucletide are done simultaneously during a 20-min incubation in the presence of a thermostable DNA ligase. After that, stringent washing with NaOH is done to remove non-ligated oligomers. Therefore, a perfect match is obtained in case of if no SNP is present in the DNA sequence and vice versa. The detector probe gets hybridized to normal DNA sequence and is detected by incubation with an antibiotin IgG-HRP conjugate due to presence of biotine. A purple precipitate is formed which can be detected by eye or a simple digital-imaging system (Fig. 2).
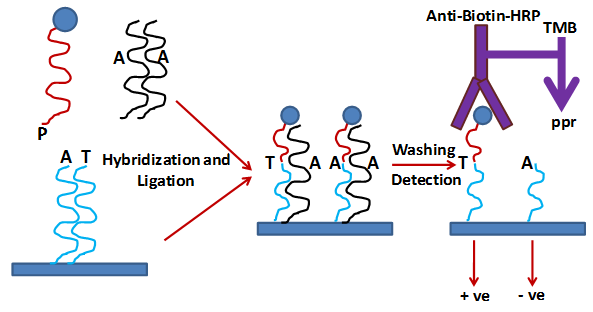
Figure 2 : Strategy for SNP detection on thin-film biosensor chips.
|