DNA Sequence Analysis Methods-II
Download this lecture
Fluorescence DNA Sequencing:
Fluorescence DNA sequencing or automated DNA sequencing implies the same procedure as in Sanger's method but with one major difference. In Sanger's method, detection is done by developing autoradiograph with radiolabelled primer which makes it lengthy and time consuming and it can be used as an ideal throughput procedure due to its health hazardousness. To get rid of these problems, in automated DNA sequencing, fluorescent molecules are being used instead of radiolabelled. Fluorescent dye labels are incorporated into DNA extension products using 5' labelled primers (dye primers) or 3' labelled dideoxynucleotide triphosphates i.e. Dye terminators. Usually the fluorescent labelling is done at 3' end of dideoxyNTP so different type of fluorochromes can be used for each ddNTP and whole reaction can be carried out in a single tube. Each dye may have a different emission wavelength when excited with an argon ion laser. Thus all four bases can be distinguished by the emission of four different colors, in a single gel lane (Fig. 5A) or by capillary electrophoresis (Fig. 5 B,C,D). Capillaries are small, a 50 μm inner diameter, and they dissipate heat very efficiently due to their high surface area to volume ratios. Thus a capillary based system can be run with much higher voltages and it dramatically lowering the run time. Most importantly, capillary systems can be automated which is a major limitation in gel based systems.
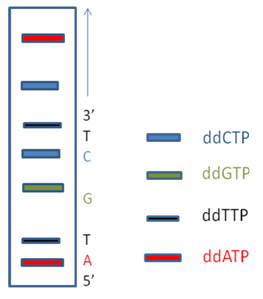
Figure 5A: A Sequencing SDS PAGE gel with 4 different fluorochrome tagged ddNTP products run in a single lane.
|